Post
Parallelize a wide df in Pandas
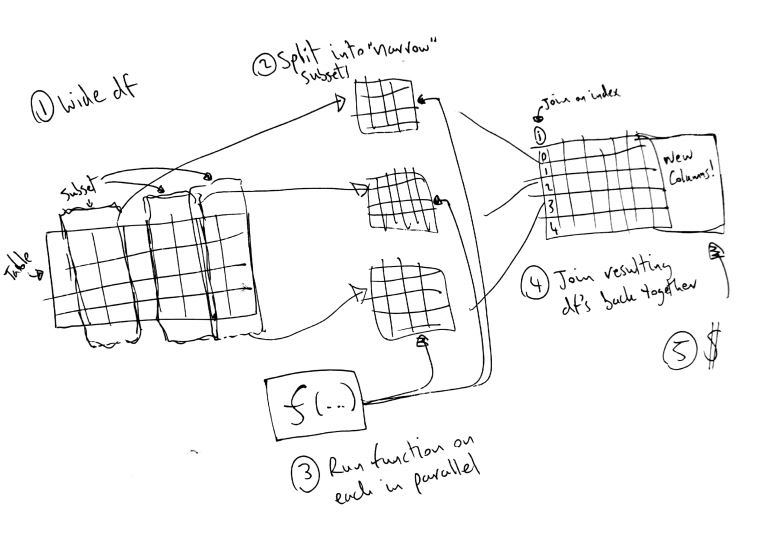
I was going to make a pretty picture.
Sometimes you end up with a very wide pandas dataframe and you are interested in doing the same types of operations (data processing, building a model etc.) but focused on subsets of the columns.
For example if we had a wide df with different time series kpi’s represented as columns then we might want to do something like look at each kpi at a time, apply some pre-processing and build something like an ARIMA time series model perhaps.
This is the situation i found myself in recently and it took me best part of an afternoon to figure out. Usually when i find myself in that situation i try and squeeze out a blog post in case might be useful for someone else or future me.
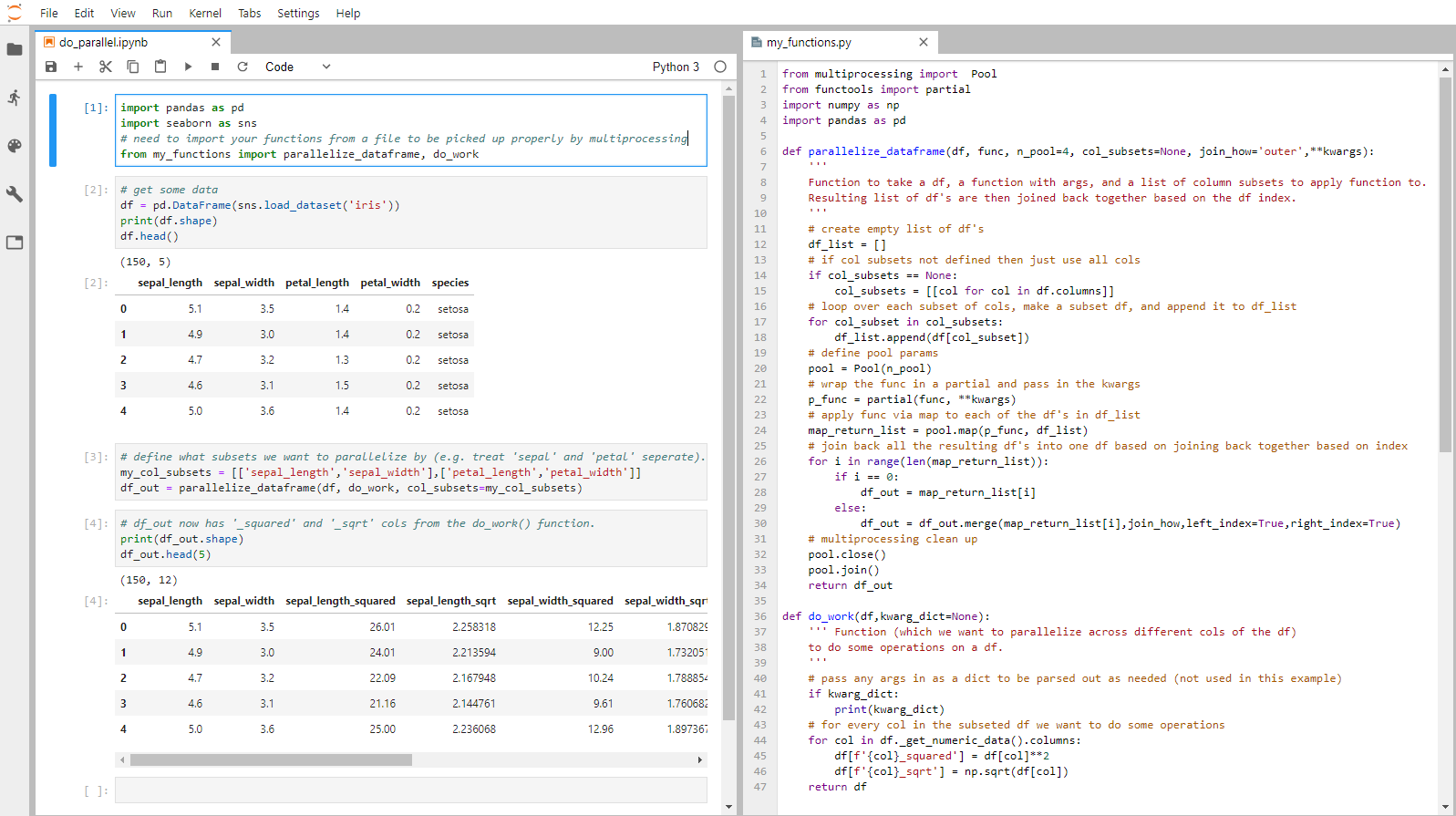
All the code in one glorious screenshot!
Note: repository with all code is here. p.s. thanks to this and this post that i built off of.
For this example i’m afraid i’m going to use the Iris dataset :0 . This example is as minimal and easy as i could throw together, basically the aim of the code is to:
- Build some function to take in a df, do some processing and spit out a new df.
- Have that function be parameterized in some way as might be needed (e.g if you wanted to do slightly different work for one subset of columns).
- Apply that function in parallel across the different subsets of your df that you want to process.
There are two main functions of interest here parallelize_dataframe() and do_work() both of which live in their own file called my_functions.py which can be imported into your jupyter notebook.
https://gist.github.com/andrewm4894/932304809840d6cef7ed5c7fc21b72cb
parallelize_dataframe() does the below things:
- Break out df into a list of df’s based on the col_subsets list passed in as a parameter.
- Wrap the function that was passed in into a partial along with the kwargs (this is how your parameters make it into the do_work() function).
- Use map() from multiprocessing to apply the func (along with the args you want to send it) to each subset of columns from your df in parallel.
- Reduce all this back into on final df by joining all the resulting df’s from the map() output into one wide df again (note the assumption here of joining back on the df indexes - they need to be stable and meaningful).
The do_work() function in this example is just a simple function to add some new columns as examples of types of pandas (or any other) goodness you might want to do. In reality in my case it would be more like a apply_model() type function that would take each subset of columns, do some feature extraction, train a model and then also score the data as needed to.
Having the ability to do this for multiple subsets of columns in your wide df can really free up your time to focus on the more important things like dickying around with model parameters and different pre-processing steps :)
That’s pretty much it, a productive afternoon (in the play center with kids i might add) and am quite pleased with myself.
Update: One addition i made to this as things got more complicated when i went to implement it was the ability to apply different function params to each subset df. For example if you wanted to pass in different parameters to the function for different columns. In the do_parallel_zip.ipynb and corresponding my_functions_zip.py (i’m calling them “_zip” as they use zip() to “zip” up both the df_list and the corresponding kwargs to go with it to be unpacked later by do_work_zip()).
To be concrete, if we wanted to multiply the “sepal_…” cols by 100 and the “petal_..” cols by 0.5. We could use the “zip” approach like below (notebook here):
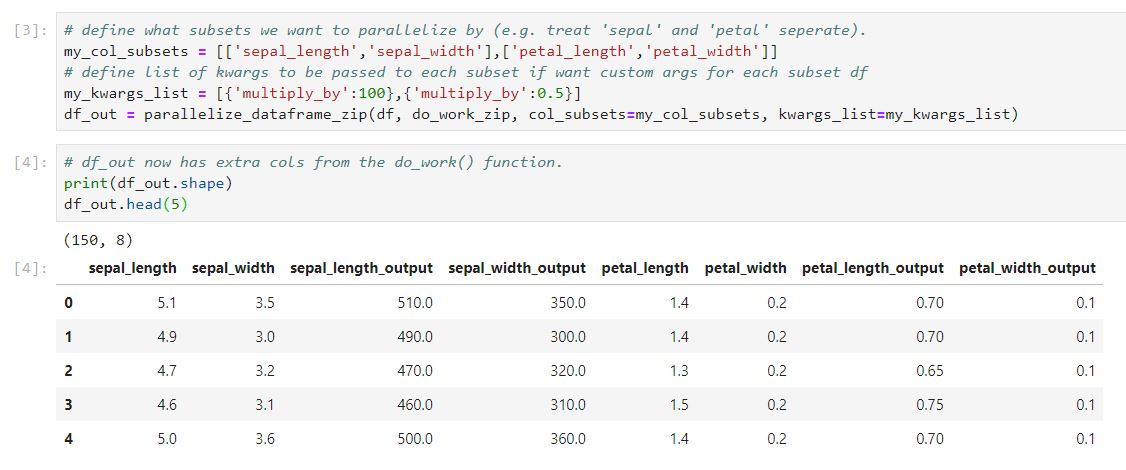
Which is using the “zip” approach in parallelize_dataframe_zip()
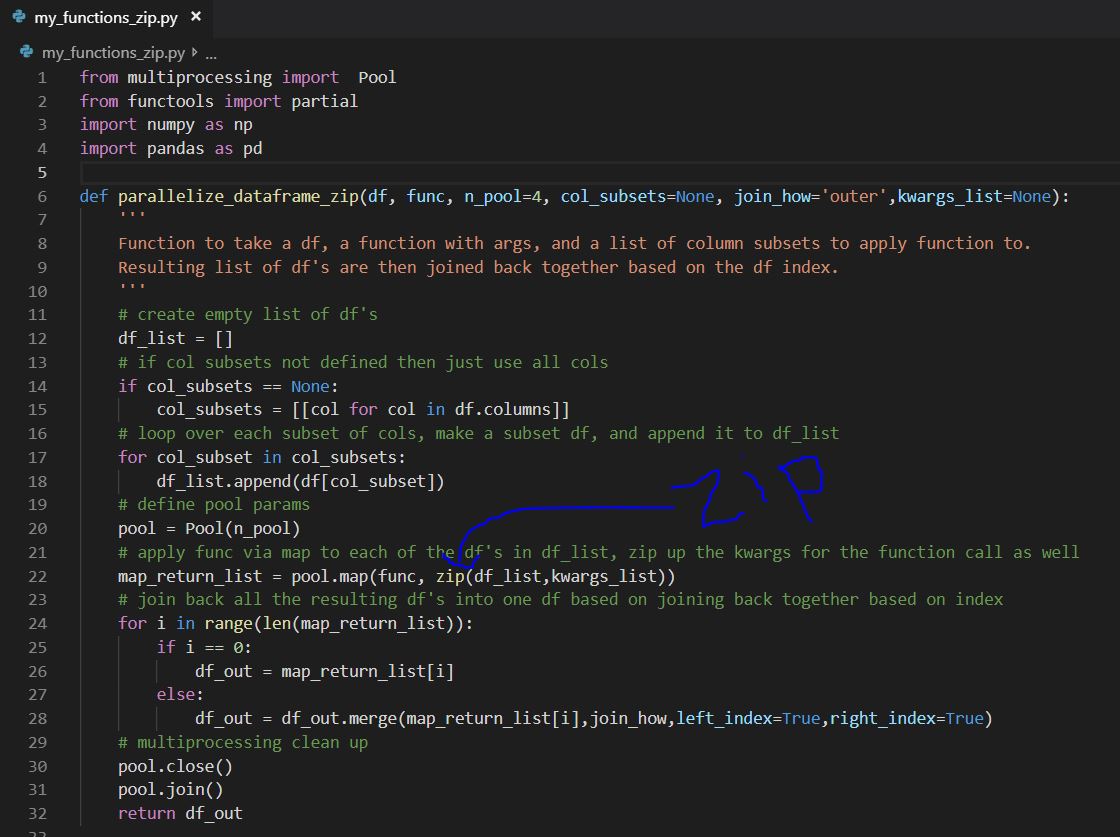
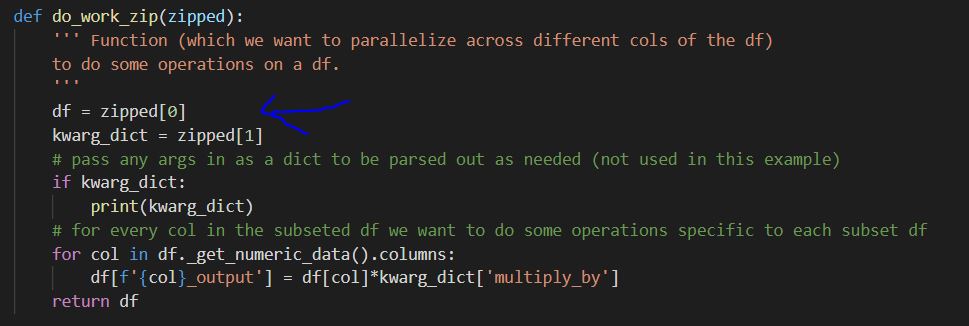
Where the zipped iterable is then unpacked as needed by the do_work_zip() function:
Comments